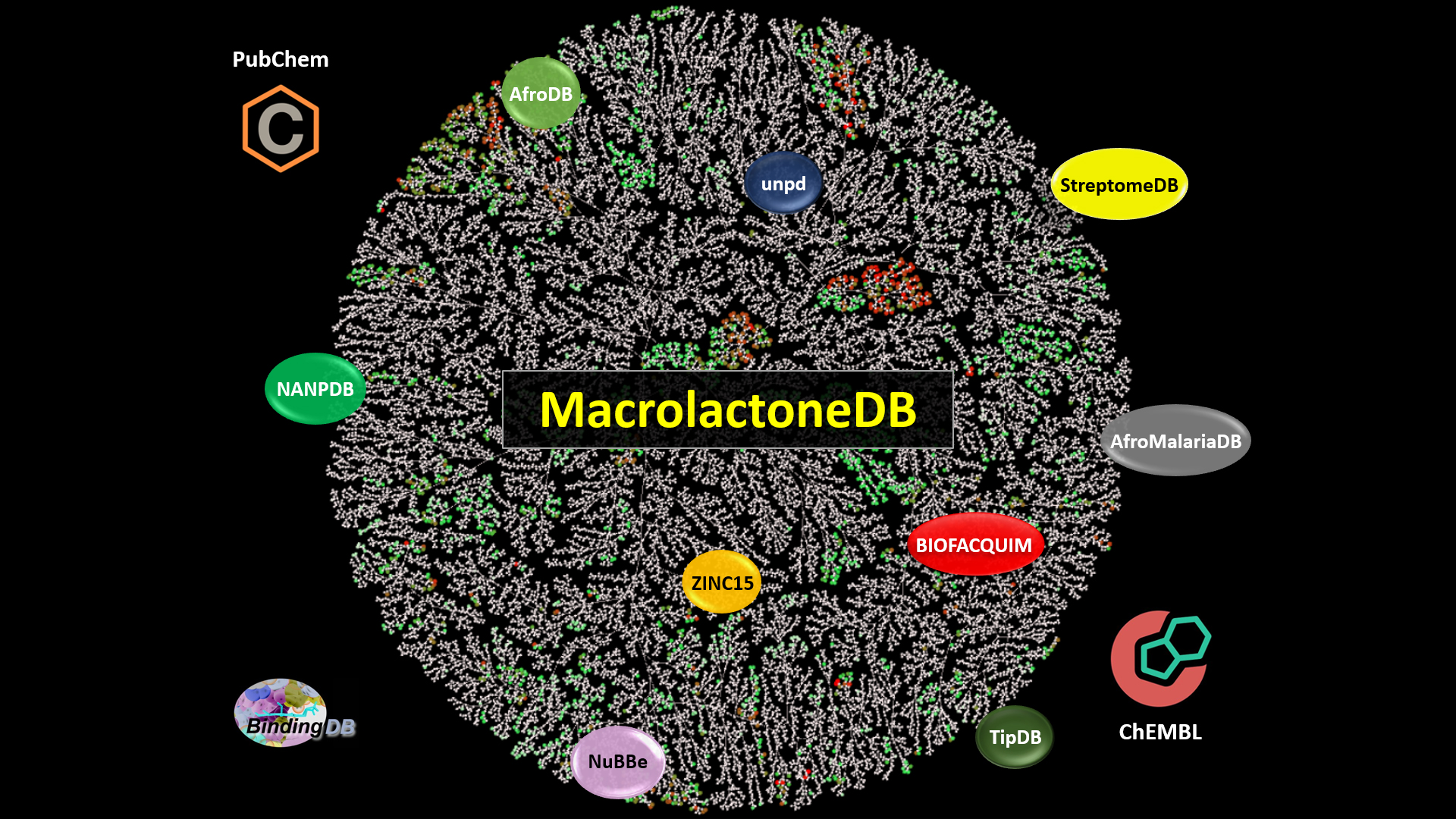
Researchers from North Carolina State University and Collaborations Pharmaceuticals have created a free-to-use database of 14,000 known macrolactones – large molecules used in drug development – which contains information about the molecular characteristics, chemical diversity and biological activities of this structural class. The database, called MacrolactoneDB, fills a knowledge gap concerning these molecules and could serve as a useful tool for future drug discovery.
Macrolactones are molecules with at least 12 atoms composing their ring-like structure. Among many useful characteristics, macrolactones’ ability to bind to difficult protein targets makes them suitable for antiviral, antibiotic, antifungal and antiparasitic drugs. However, their size and complicated structure make them difficult to synthesize.
“Macrolactones are titanic molecules – their size presents challenges to researchers who may want to work with them,” says Sean Ekins, CEO of Collaborations Pharmaceuticals, member of NC State’s Comparative Medicine Institute, entrepreneur in residence at UNC-Chapel Hill’s Eshelman School of Pharmacy and corresponding author of the research. “We wanted to address that issue by creating a publicly available database of these molecules and their properties.”
NC State graduate student and first author of the paper Phyo Phyo Zin mined 13 public databases for 14,000 known macrolactones, compiling them into MacrolactoneDB. Only 20% of the macrolactone compounds she curated had biological data associated with them.
Zin, Ekins and NC State Associate Professor of Chemistry Gavin Williams conducted cheminformatics analyses of the macrolactones’ molecular properties and developed 91 descriptors to better characterize the molecules. The researchers then looked at three targets of interest for some of the macrolactones – specifically malaria, hepatitis C and T cells – and used machine-learning techniques to understand the structure-activity relationship between the macrolactones and these targets.
“We know that macrolactone drugs are effective, but there’s a lot we don’t know about what makes a good one,” Williams says. “That’s why we set out to do this research. We found that it is possible to utilize machine learning with these molecules, and improving our analysis and description of macrolactones will improve prediction models going forward.”
“Anyone interested in these molecules or in drug development utilizing macrolactones now has a user-friendly database where everything is accessible and in one location,” Ekins says. “Researchers can ask questions about what makes a particular macrolactone molecule well-suited for a particular biological application.
“Hopefully MacrolactoneDB will help us to understand this diverse class of molecules, and move forward in creating new ones.”
The work appears in Scientific Reports and was supported by the National Institutes of Health under grants R44GM122196-02A1 and R43AT010585-01S1. Zin received additional funding from the American Association of University Women and an NC State Graduate Research Assistantship.
-peake-
Note to editors: An abstract follows.
“Cheminformatics Analysis and Modeling with MacrolactoneDB”
DOI: 10.1038/s41598-020-63192-4
Authors: Phyo Phyo Kyaw Zin, Gavin Williams, Sean Ekins, North Carolina State University; Sean Ekins, Collaborations Pharmaceuticals, Inc.
Published: April 15, 2020 in Scientific Reports
Abstract:
Macrolactones, macrocyclic lactones with at least twelve atoms within the core
ring, include diverse natural products such as macrolides with potent
bioactivities (e.g. antibiotics) and useful druglike characteristics. We have
developed MacrolactoneDB, which integrates nearly 14,000 existing macrolactones
and their bioactivity information from different public databases, and new
molecular descriptors to better characterize macrolide structures. The chemical
distribution of MacrolactoneDB was analyzed in terms of important molecular
properties and we have utilized three targets of interest (Plasmodium
falciparum, Hepatitis C virus and T-cells) to demonstrate the value of
compiling this data. Regression machine learning models were generated to
predict biological endpoints using seven molecular descriptor sets and eight
machine learning algorithms. Our results show that merging descriptors yields
the best predictive power with Random Forest models, often boosted by consensus
or hybrid modeling approaches. Our study provides cheminformatics insights into
this privileged, underexplored structural class of compounds with high
therapeutic potential.
This post was originally published in NC State News.
- Categories: